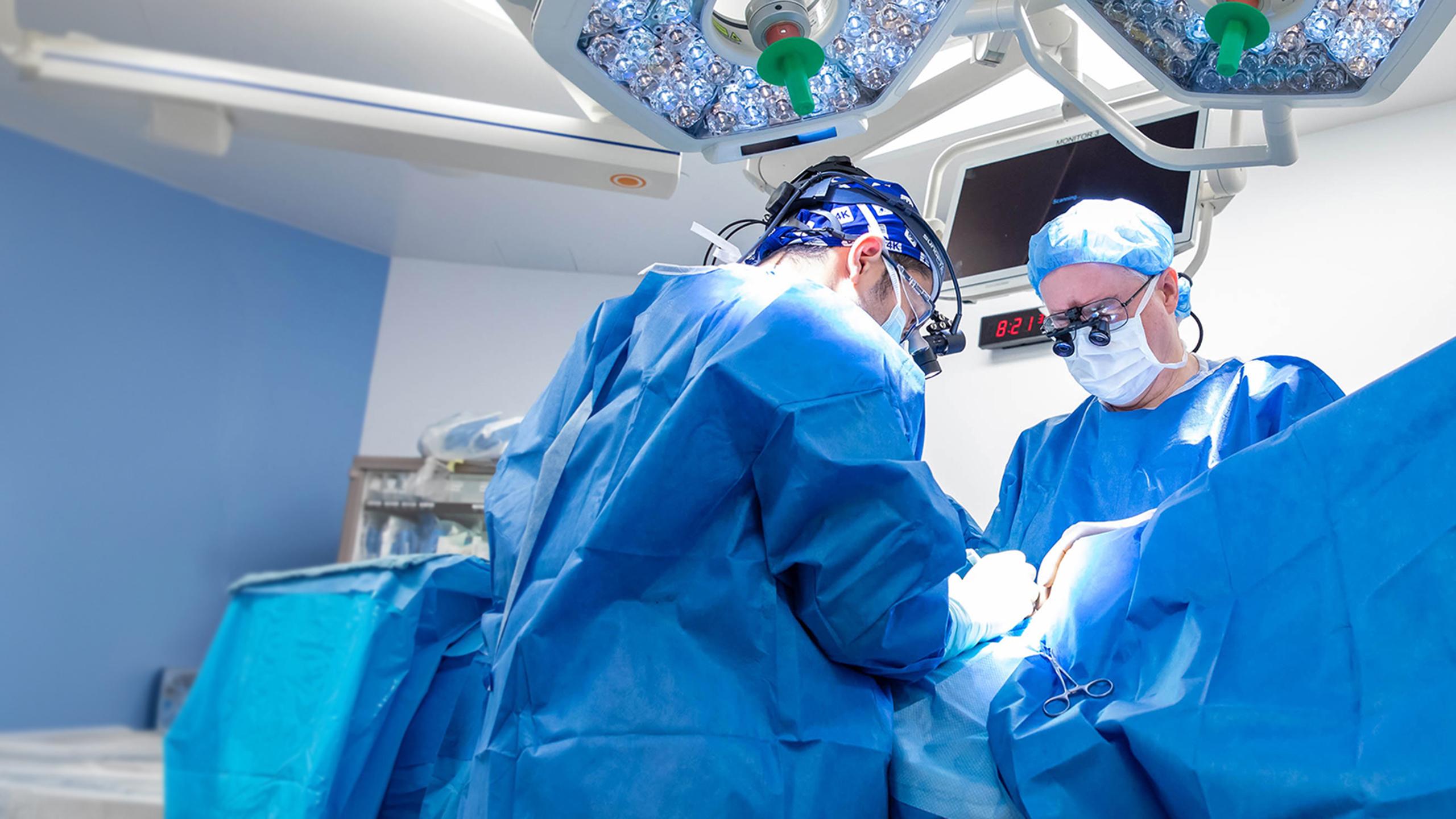
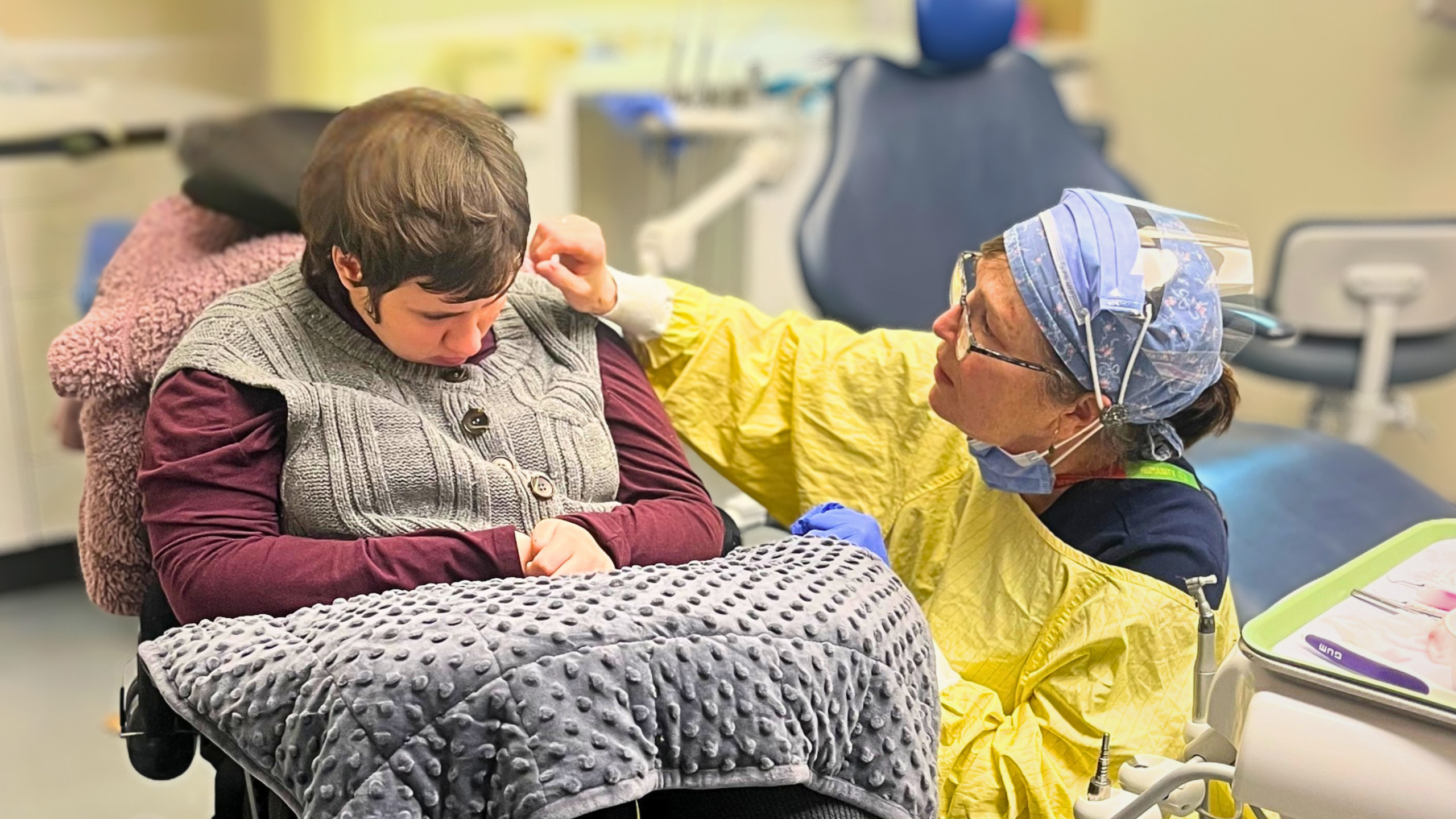
Dr. Daniel Schramek
Lunenfeld-Tanenbaum Research Institute
Deputy Director, Discovery Research, Sinai Health
Functional genomics for precision oncology
Our lab is dedicated to revolutionizing cancer treatment through functional genomics, aiming for personalized and highly specific therapies. We focus on identifying novel treatments for the most aggressive cancers by systematically uncovering genotype-specific vulnerabilities – the unique weaknesses of cancer cells based on their genetic makeup.
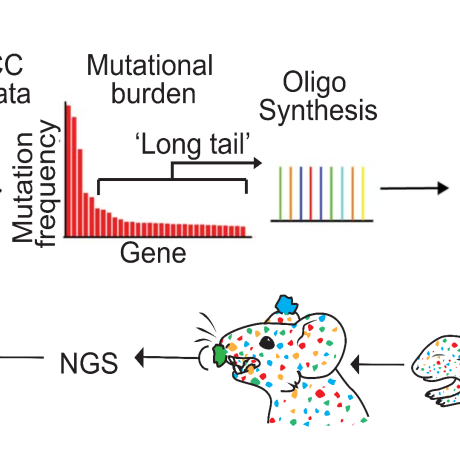
A cornerstone of our research is the development and application of cutting-edge gene-editing approaches in animal models. This innovative approach allows us to seamlessly integrate complex human cancer genomics data with sophisticated mouse modelling. By doing so, we can functionally assess the roles of hundreds of putative cancer genes, evaluating their capacity to drive tumour formation (tumourigenesis) and spread (metastasis) directly within a living organism. This approach takes into account the critical interactions between cancer cells and their environment that cannot be replicated in a laboratory dish.
This powerful combination of high-throughput screening, advanced gene-editing technology and relevant in vivo models enables us to rapidly pinpoint critical cancer drivers and dependencies. Ultimately, our goal is to translate these fundamental discoveries into new therapeutic strategies, paving the way for more effective and tailored treatments for patients battling devastating malignancies. We believe this direct functional assessment in a physiological context is key to accelerating the discovery of impactful cancer therapies.
Room 1084, 600 University Avenue
Toronto, M5G 1X5
Website: Schramek Lab
Publications: Pubmed
Google Scholar: Daniel Schramek
ORCID: 0000-0001-9977-2104
LinkedIn: Daniel Schramek
- 2025–present; Professor, Department of Molecular Genetics, Temerty Faculty of Medicine, University of Toronto, Toronto
- 2024–present; Deputy Director for Discovery Research and Tony Pawson Chair, Sinai Health, Toronto
- 2022–present; Senior Investigator, Lunenfeld-Tanenbaum Research Institute, Sinai Health, Toronto
Former appointments
- 2022–2025; Associate Professor, Department of Molecular Genetics, Temerty Faculty of Medicine, University of Toronto, Toronto
- 2015–2022; Investigator, Lunenfeld-Tanenbaum Research Institute, Sinai Health, Toronto
- 2015–2022; Assistant Professor, Department of Molecular Genetics, Temerty Faculty of Medicine, University of Toronto, Toronto
- MSc in Technology Management, New York University, New York, NY, USA; 2012–2014
- Postdoctoral Fellowship & Emerald Foundation Young Investigator with Dr. Elaine Fuchs, The Rockefeller University & Howard Hughes Medical Institute, New York, NY, USA; 2011–2015
- PhD, University of Vienna, Vienna, Austria; 2006–2010
- MSc, University of Vienna, Vienna, Austria; 2000–2006
- 2024 – Tony Pawson Sinai 100 Chair
- 2021 – Canadian Research Chair in Functional Cancer Genomics, tier II (renewed)
- 2020 – ASCINA Award; Austria
- 2016 – Human Frontier of Science Program (HFSP) Career Development Award
- 2016 – Early Research Award (ERA), Ontario; Canada
- 2016 – Susan Komen Career Catalyst Award
- 2016 – Canadian Research Chair in Functional Cancer Genomics, tier II
- 2015 – Venture Sinai Scientist Award, Mount Sinai Hospital Foundation, Canada
- 2015 – CCSDP New PI Award, CIHR-ICR and CCSRI; Canada
- 2014 – Regeneron Award for Creative Innovation
- 2014 – Keystone Future of Science Fund Award, Keystone Symposia; USA
- 2012 – Human Frontier Science Program (HFSP) long-term postdoctoral fellowship
- 2012 – Helen Hay Whitney Postdoctoral Fellowship, New York; USA – declined
- 2012 – Austrian Program for Advanced Research and Technology `APART´ Fellowship
- 2011–2014 – Young Investigator Award, Emerald Foundation, New York; USA
- 2011 – American Association for Cancer Research (AACR) Anna D. Barker Fellowship
Decoding aneuploidy: identifying driver genes and therapeutic targets in recurrent CNAs
The prevalent issue of recurrent Copy Number Alterations (CNAs) across devastating malignancies including breast, lung, brain, HNSCC, and pancreatic cancers is a critical challenge in cancer therapy. While these large-scale genomic gains and losses are common, identifying the specific "driver" genes within these altered regions that fuel cancer progression, and the unique vulnerabilities they create, remains a significant hurdle.
Guided by principles similar to those demonstrated in recent cutting-edge in vivo functional genomics studies, we are employing advanced in vivo gene editing and screening technologies such as CRISPR-KOALA. Our strategy involves systematically dissecting these complex CNA regions directly within physiological models. This allows us to functionally test the contribution of numerous candidate genes located within these amplified or deleted segments.
By doing so, we aim to distinguish true oncogenic drivers from mere passengers and, crucially, to uncover novel, context-specific therapeutic vulnerabilities associated with these recurrent CNAs. This approach allows for a high-throughput, physiologically relevant assessment, moving beyond correlative genomic data to establish causal links. Ultimately, this project seeks to pinpoint robust therapeutic targets within these challenging genomic alterations, paving the way for innovative and more effective targeted treatments for patients suffering from these aggressive cancers.
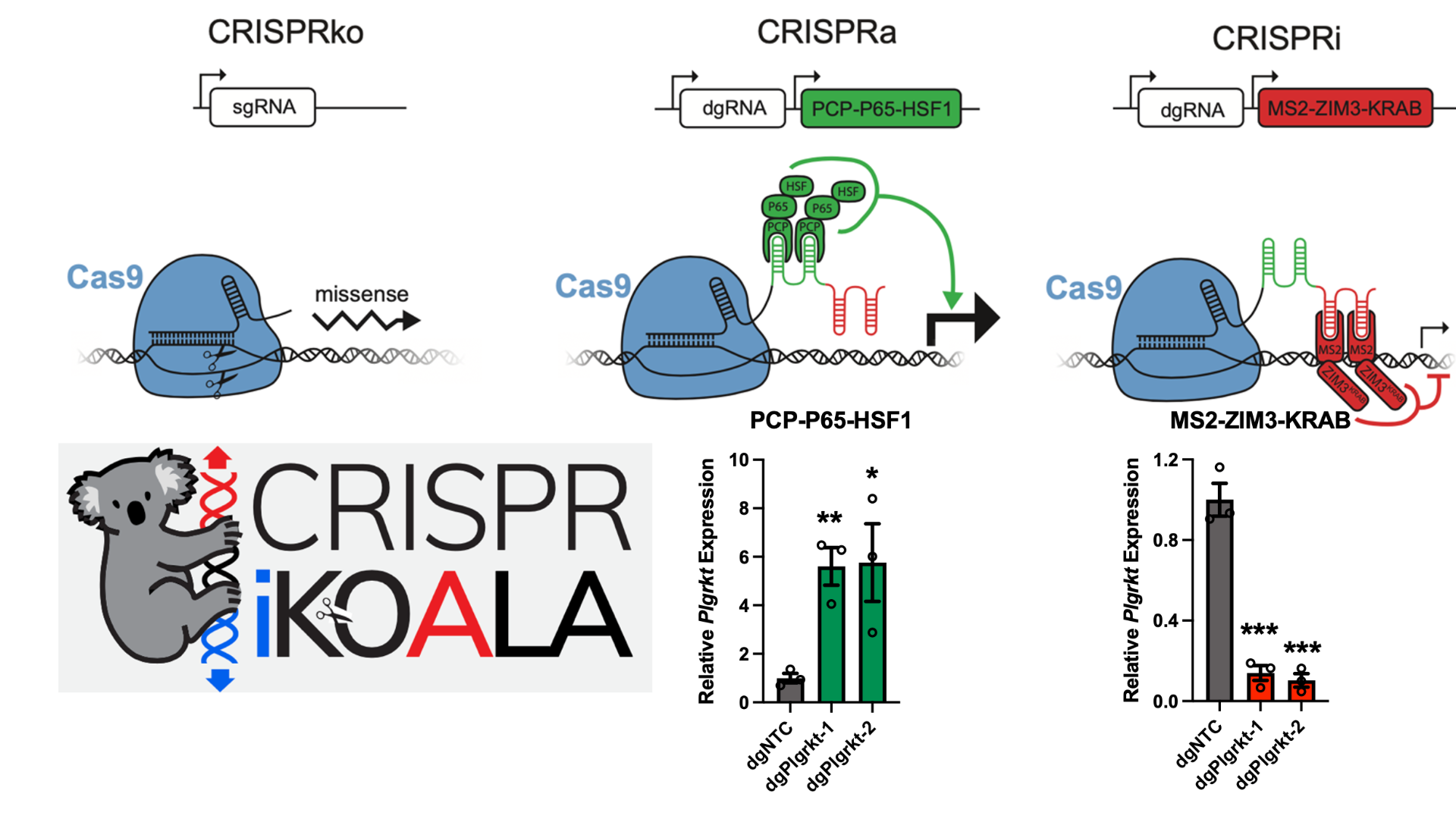
Elucidating the landscape of essential genes in various epithelial tissues and under specific oncogenic alterations
Our research is pioneering new frontiers in identifying cancer's Achilles' heels through the development and application of our novel CRISPR-StAR2.0 technology. This cutting-edge platform is specifically engineered to uncover oncogene-specific vulnerabilities - critical dependencies that cancers develop based on the particular oncogenes driving their growth - across a spectrum of malignancies. A major challenge in targeted therapy is understanding how different oncogenes sculpt unique vulnerabilities in diverse tissue environments.
The CRISPR-StAR2.0 system allows us to conduct highly sophisticated functional genomic screens directly in vivo, within physiologically relevant mouse models that accurately recapitulate human cancers in various organs. This powerful approach enables us to systematically perturb genes in the presence of specific oncogenic drivers, precisely mapping out the genetic interactions and dependencies that are essential for tumor survival and progression in a given context. By interrogating these complex biological networks directly in living systems, CRISPR-StAR2.0 overcomes limitations of traditional in vitro screens.
Our goal is to generate a comprehensive atlas of actionable therapeutic targets that are uniquely tailored to cancers driven by distinct oncogenes. This project promises to significantly accelerate the discovery of next-generation precision medicines, offering new hope by exploiting the specific molecular wiring dictated by the oncogenes at the heart of each cancer.
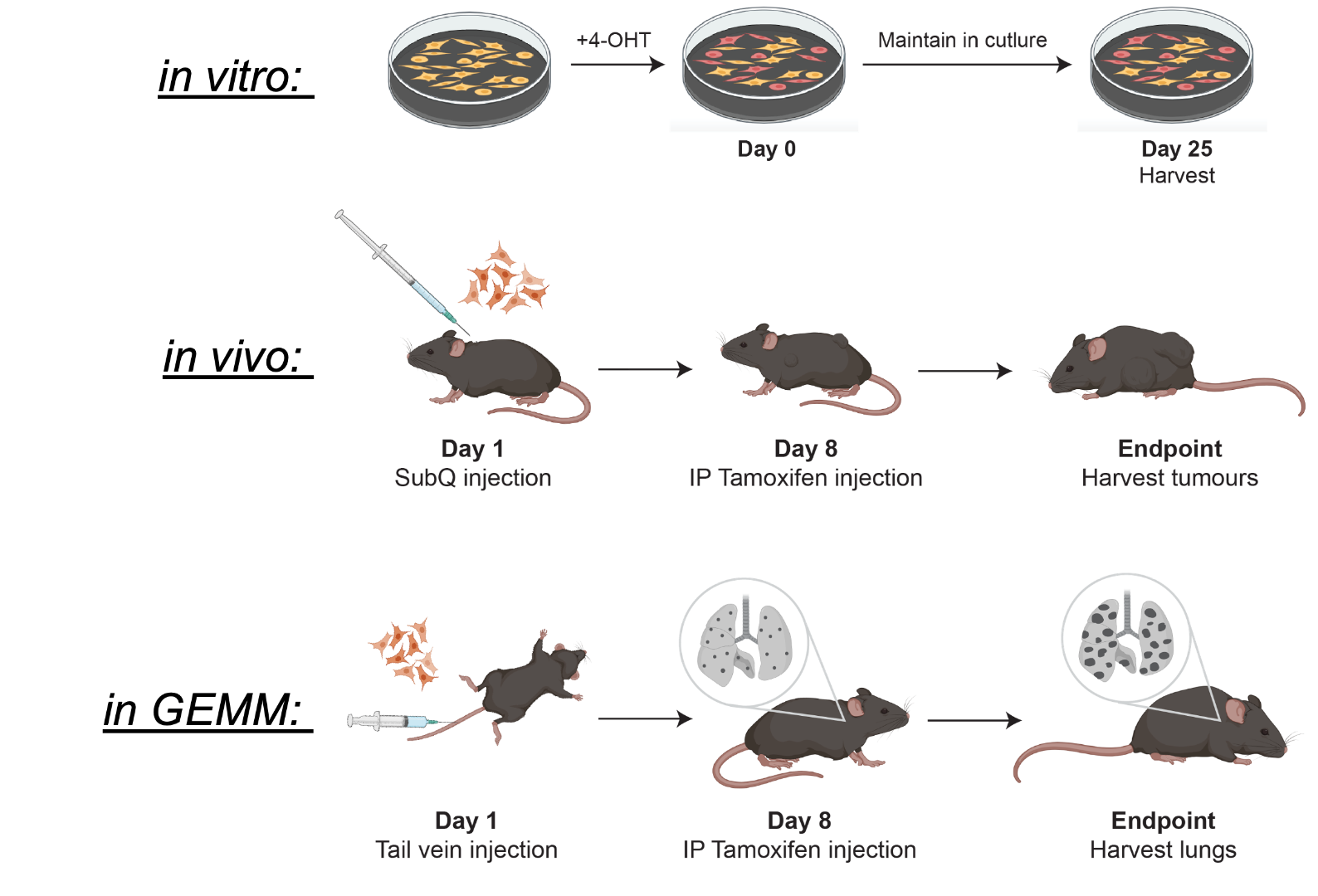
We are always looking for motivated researchers to join our team.
Postdocs
Our research group is always interested in recruiting highly motivated postdoctoral fellows with a strong publication record in cancer biology. Please forward your CV, references and research interests to [email protected].
Graduate students
Our research group is part of the Department of Molecular Genetics at U of T, which has a central admission committee and a rotation system. Graduate students interested in doing a PhD in the laboratory must first be accepted in the Department of Molecular Genetics at U of T.
Notable publications
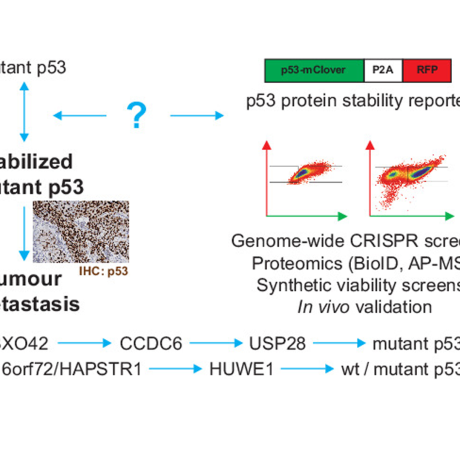
Molecular Systems Biology, 2024
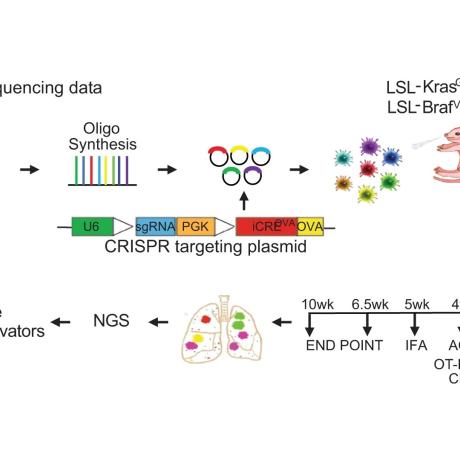
Nature Communications, 2023
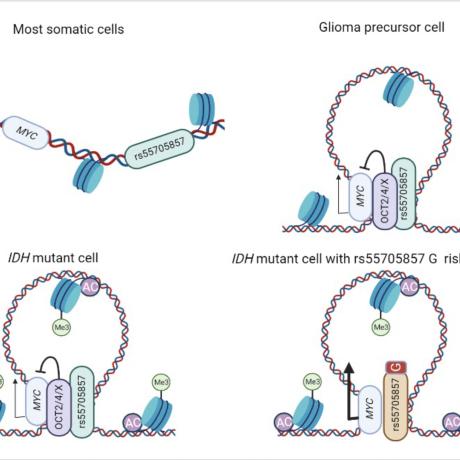
Science, 2022
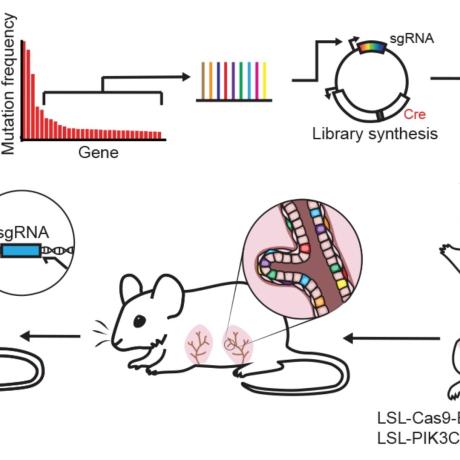
Cancer Discovery, 2022
Science, 2020
Join our team
Visit our job board to see research positions.