Dr. Alissa Greenwald
Lunenfeld-Tanenbaum Research Institute
Turning the spatial logic of cancer into clinical opportunity
We are a cancer systems biology lab investigating the organizational principles of tumours with the goal of therapeutically exploiting these principles to develop better cancer treatments. We profile patient tumour samples to generate detailed spatial gene expression maps. Understanding which cell states and cell types tend to consistently associate with each other and how they interact within a community can be leveraged as a therapeutic vulnerability.
To gain a holistic view of tumour organization, we are developing experimental and computational approaches to spatial biology that identify the most robust patterns and interactions across molecular, cellular and tissue scales, test their essentiality and the functional consequences of perturbing them.
We study tumour organization through two different approaches. First, we spatially profile patient tumour samples and use quantitative, systems biology approaches to identify the most consistent and robust patterns across tumours. Additionally, we ask how these patterns relate to the genetics and clinical features of the tumour. Second, we ask how tumours are built using model systems to impair or promote organization. Collectively, these approaches can help us to understand how to therapeutically modulate tumour organization.
Email: [email protected]
Room 989, 600 University Avenue
Toronto, M5G 1X5
Website: Greenwald Lab
Publications: PubMed
Google Scholar: Alissa Greenwald
ORCID number: 0000-0001-5428-9645
Discovery Research: Alissa Greenwald
Bluesky: Alissa Greenwald
X: Alissa Greenwald
- 2025–present; Investigator, Lunenfeld-Tanenbaum Research Institute, Sinai Health, Toronto
- 2025–present; Assistant Professor, Department of Molecular Genetics, University of Toronto, Toronto
- Postdoctoral fellow and staff scientist in Computational Systems Biology with Dr. Itay Tirosh, Weizmann Institute of Science, Rehovot, Israel; 2018–2025
- PhD in Developmental Biology and Cancer Research with Dr. Eli Keshet, Hebrew University of Jerusalem, Jerusalem, Israel; 2012–2017
- MSc in Biomedical Sciences, Hebrew University of Jerusalem, Jerusalem, Israel; 2009–2011
- BA in Biological Chemistry, Wellesley College, Wellesley, Massachusetts, USA; 2002–2006
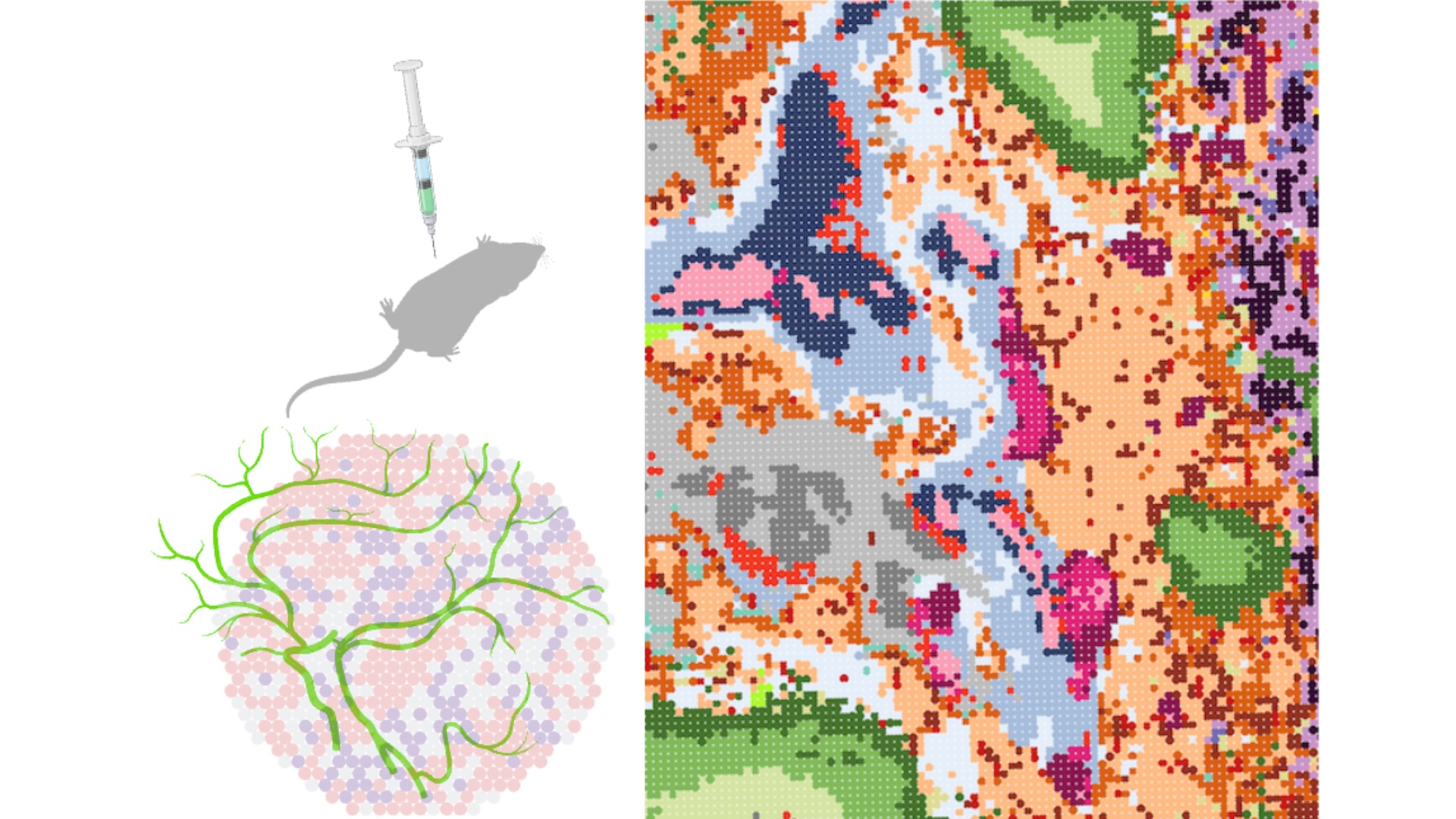
Mapping of patient tumours towards spatially-aware medicine
With the advent of spatial-omics, our ability to quantitatively map tumour organization in situ at the level of cell states, cell types and genetic subclones has improved dramatically. We use detailed tumour spatial maps to identify patterns that repeat robustly across tumours. Knowing which cell states and cell types tend to associate with each other and how they interact within a community can be leveraged as a therapeutic vulnerability. We integrate spatial information across molecular, cellular and tissue scales in order to understand the principles of tumour organization. Our current focus is on tumours of the central nervous system (CNS) and sarcomas.
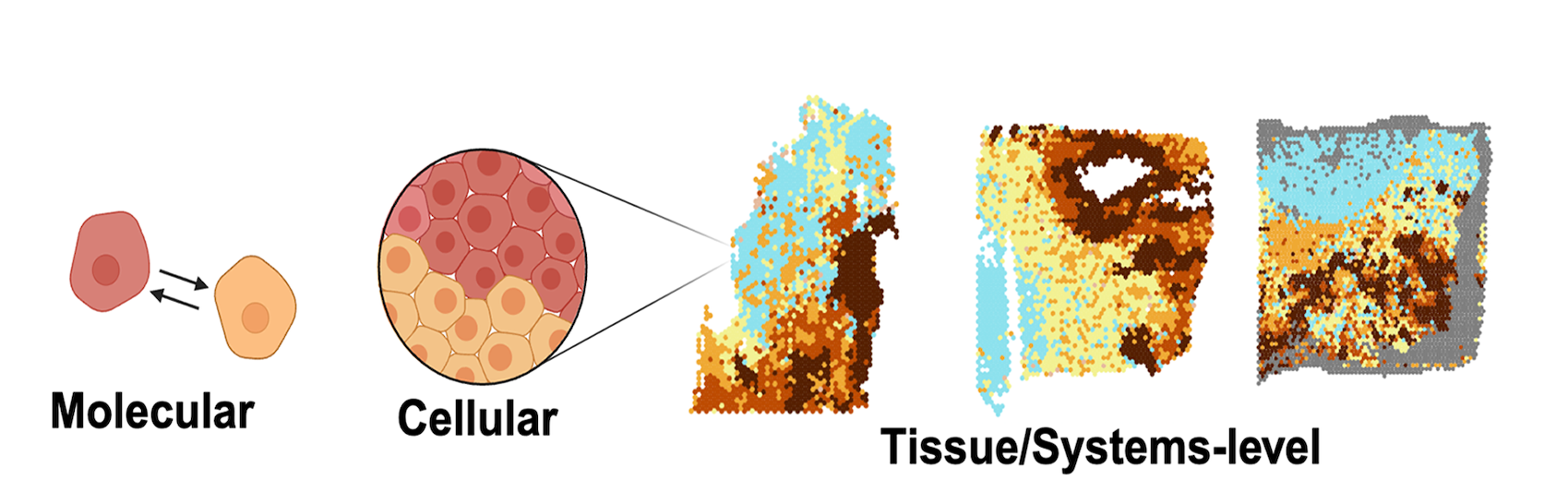
Do tumours utilize developmental mechanisms of tissue organization and spatial patterning?
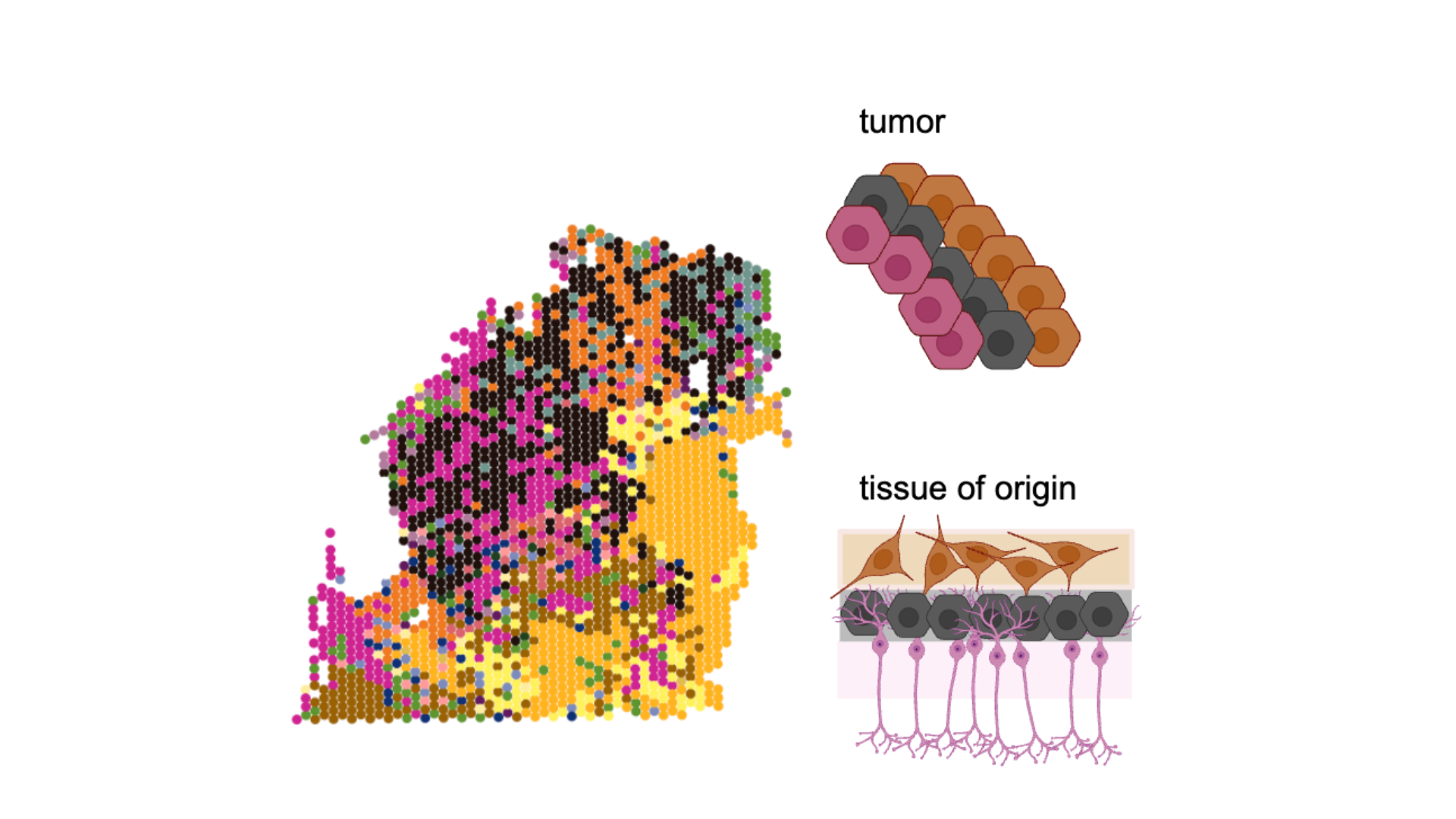
Cancer cells often mimic partial or aberrant versions of normal developmental expression programs such as epithelial-to-mesenchymal transition or senescence. Our recent work suggests that in some contexts, these cancer cell states also mimic spatial associations observed in their tissue of origin.
In addition to looking for patterns that repeat between tumours we can also look for patterns that repeat between a tumour and its tissue of origin in the adult or during development. On one hand, a hallmark of cancer is a lack of responsiveness to environmental signals; on the other hand, even in advanced, aggressive disease, tumours can exhibit spatial patterning suggestive of responsiveness to long-range environmental cues.
We are exploring this paradox in order to better understand the dichotomy of structure vs. disorganization in cancer and how it relates to tumour evolution and plasticity.
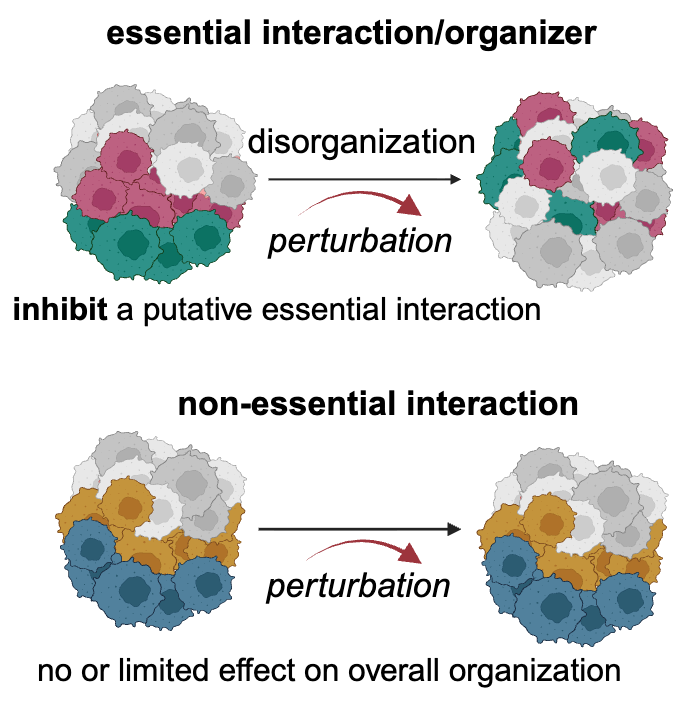
Modeling and perturbing tumour organization
Development of integrative spatial profiling technologies
We are seeking curiosity-driven and kind scientists at all levels with interests in combining computational and experimental biology to address fundamental questions in tumor organization. We aim to build an interdisciplinary, open-minded, and inclusive lab community.
Postdoctoral fellows
Please email [email protected] your CV and a description of your research interests.
Graduate students
Our research group is part of the Department of Molecular Genetics at the University of Toronto, which has a central admission committee and a rotation system. Graduate students interested in doing a PhD in the laboratory must first be accepted to the Department of Molecular Genetics.
Summer students
Summer students are exclusively selected from successful applicants to the Research Training Center (RTC) at the Lunenfeld-Tanenbaum Research Institute. Applications are available online and need to be completed by February 28th of each year.
Notable publications
Nature Communications, 2026
Cell, 2024
Nature, 2023
Nature Genetics, 2020
Join our team
Visit our job board to see research positions.