Dr. Williams Turpin
Lunenfeld-Tanenbaum Research Institute
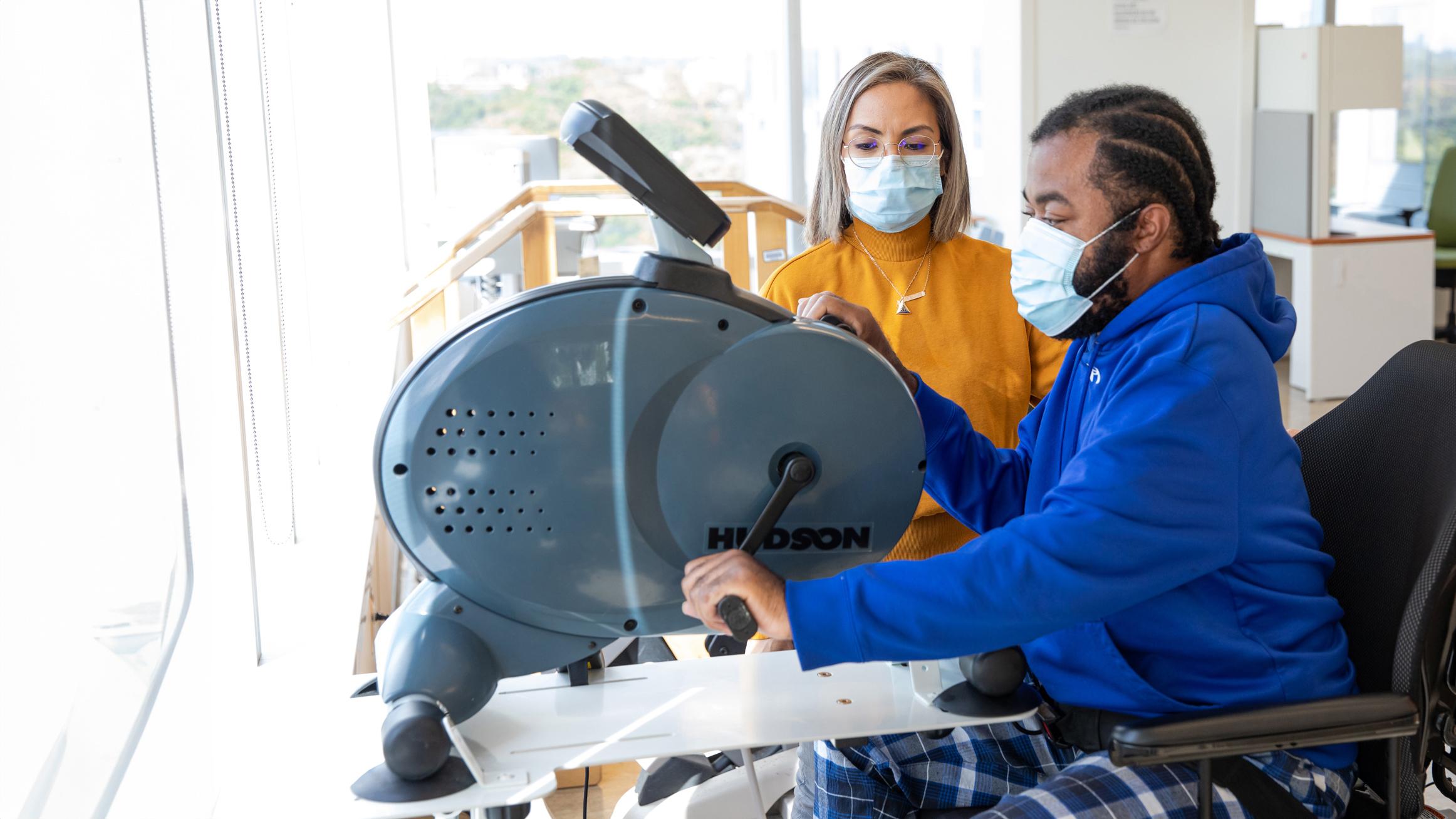
Our research looks at how what we eat and the bacteria living in our gut influence the risk of developing Crohn’s disease, a type of inflammatory bowel disease (IBD). Crohn’s disease usually starts in young adults and has no cure, so finding ways to prevent it is very important.
We study large groups of people who are at risk of Crohn’s disease and combine this with advanced tools that measure gut bacteria, diet, and signals from the body. This helps us understand which foods and bacterial activities protect the gut and which ones may trigger inflammation.
The goal of our work is to identify everyday factors—like diet and gut health—that we can change to lower the risk of Crohn’s disease. By learning how gut microbes and diet work together, we hope to design simple prevention strategies, such as nutrition guidance or safe bacteria-based therapies, that can help people stay healthy.
This research matters because it focuses on prevention and improving quality of life, with the potential to reduce the burden of Crohn’s disease for patients, families, and the healthcare system.
Email: [email protected]
Room L3-012, 60 Murray Street
Toronto, M5T 3L9
Publications: PubMed
Google Scholar: Williams Turpin
ORCID: 0000-0001-9364-5868
U of T Profile: Williams Turpin
LinkedIn: Williams Turpin
- 2025–current; Assistant Professor, Department of Nutritional Science, University of Toronto, Toronto
- 2025–current; Translational Research Scientist, Lunenfeld-Tanenbaum Research Institute, Sinai Health, Toronto
Former appointments
- 2020–2024; Senior Research Associate, Department of Medicine, University of Toronto, Toronto
- 2017–2020; Research Associate, Department of Medicine, University of Toronto, Toronto
- Postdoctoral Fellowship, University of Toronto, Toronto; 2012–2017
- PhD in Food Science, Université Montpellier II, Montpellier, France; 2006–2011
- Université Versailles / St Quentin, France
- Digestive Disease Week 2016 – Young Investigator Award for the abstract in Microbiome: Structure and Function, San Diego, California, United States for the abstract “Ileal pouch symptoms are associated with changes in microbial composition and diversity not necessarily related to endoscopic inflammation”.
- United European Gastroenterology Week 2015 – National Scholar Award for the best Canadian abstract entitled “Genetic influence on composition of the intestinal bacteria in healthy first degree relatives of Crohn’s disease subjects”. United European Gastroenterology week 2015 (24-28 October 2015), Barcelona, Spain.
- Digestive Disease Week 2015 – Young Investigator Award for the distinguished abstract in plenary session, IMIBD Distinguished Abstract Plenary session, Digestive Diseases Week 2015 (16-19 May 2015) Washington, District of Columbia, United States for the abstract “Genome-wide association of composition and diversity of the intestinal bacterial phyla in healthy first degree relatives of Crohn’s Disease subjects”.
Toward personalised nutritional approach to prevent or manage inflammatory bowel disease
Crohn’s disease (CD) is a chronic inflammatory bowel condition with rising incidence worldwide. While genetics and environment contribute, diet is a key modifiable risk factor. Large prospective studies show that high fiber, fruit, vegetable, and fish intake lower CD risk, while Western dietary patterns rich in processed foods and saturated fats increase risk. However, inconsistent results across studies suggest that individuals respond differently to diet depending on their microbiome and physiology.
The POP 2021 project aims to design personalized diets that reduce subclinical inflammation and lower the risk of CD onset. Building on GEM’s microbiome-based risk score (GMRS) and fecal calprotectin (FCP) as biomarkers of risk, this study will recruit first-degree relatives of CD patients with elevated FCP. Participants will undergo a 7-week dietary intervention alternating between Western and Mediterranean-style diets, while providing daily stool samples and food records. Microbiome sequencing and biomarker analysis will assess individual responses.
Using a Bayesian machine learning framework, the project will generate “n-of-1” personalized dietary prediction models to identify specific foods that improve inflammation and microbiome risk scores. These models will guide tailored dietary recommendations and also inform global patterns of protective food choices for at-risk populations.
This pilot study represents the first attempt to prevent Crohn’s disease through personalized nutrition. If successful, it will lay the foundation for larger trials and cost-effective, non-pharmacological strategies to reduce CD incidence globally.
Culturing and screening for gut bacteria with beneficial properties to prevent or manage inflammatory bowel disease
Crohn’s disease (CD) is a chronic inflammatory bowel disease with no cure, rising prevalence, and high personal and societal costs. Preventative strategies are urgently needed, particularly those targeting the gut microbiome, which plays a central role in CD pathogenesis. Sinai Health’s GEM Project, a prospective cohort of 5,000 healthy first-degree relatives (FDRs) of CD patients, has identified microbial signatures and metabolites associated with protection from disease onset. Building on this foundation, the proposed project seeks to isolate, characterize, and validate gut bacterial strains with therapeutic potential.
The central hypothesis is that specific strains enriched in genetically predisposed but disease-free individuals exert protective effects via immunomodulation and production of anti-inflammatory metabolites such as eugenol, short-chain fatty acids (SCFAs), and N-acylated derivatives. Three specific aims will guide the research: 1) Culturing and identifying protective bacterial strains, particularly within the Ruminococcus and Oscillibacter genera, from GEM participants who remained healthy for over a decade;
2) Functional characterization by screening isolates for safety, metabolite production, and capacity to modulate immune pathways (e.g., suppressing NF-κB activation, inducing IL-10); 3) Testing prioritized strains in the T-cell transfer colitis model, a robust system for chronic immune-mediated inflammation, to establish efficacy in reducing intestinal inflammation and enhancing barrier integrity.
This work will generate a prioritized library of candidate live biotherapeutic products, mechanistic insights into host–microbe interactions, and intellectual property opportunities. Ultimately, it lays the foundation for microbiome-based interventions to prevent or treat CD, representing a transformative, non-pharmacological approach with broad translational potential.
Predicting risk of IBD before it starts
Inflammatory bowel disease (IBD), including Crohn’s disease and ulcerative colitis, often develops silently for years before symptoms appear. Right now, there is no way to identify who is at highest risk, which means prevention is not possible. The project aims to change this by creating a simple blood test that can predict the risk of developing IBD before it begins.
This project uses samples from three unique groups: healthy relatives of Crohn’s disease patients (the GEM Project), U.S. military service members (the PREDICT cohort), and participants in the Nurses’ Health Study. All of these groups provided blood samples years before some individuals later developed IBD.
By analyzing these samples with advanced tools that measure proteins, metabolites, sugars, and immune responses, researchers will look for biological “signatures” that predict disease. The goal is to develop the first validated test to identify people at risk and ultimately guide early prevention strategies.
Our research group is part of the Department of Nutritional Science at the University of Toronto, which requires that a supervisor be identified before admission to the graduate program. Graduate students interested in doing a PhD in the laboratory/group should first contact Dr. Williams Turpin ([email protected]) directly.
Summer students are exclusively selected from successful applicants to the Research Training Center (RTC) at the Lunenfeld-Tanenbaum Research Institute. Applications are available online and need to be filled by February 28th of each year.
Notable publications
Gastroenterology, 2023
BMJ Gut, 2023
Gastroenterology, 2022
Gastroenterology, 2020
Nature Genetics, 2016
Join our team
Visit our job board to see research positions.