Sinai Health scientists provide detailed map to understanding human cells

Researchers from Sinai Health have published a study providing an ultra-detailed look at the organization of a living human cell, providing a new tool that can help scientists around the world better understand what happens during disease.
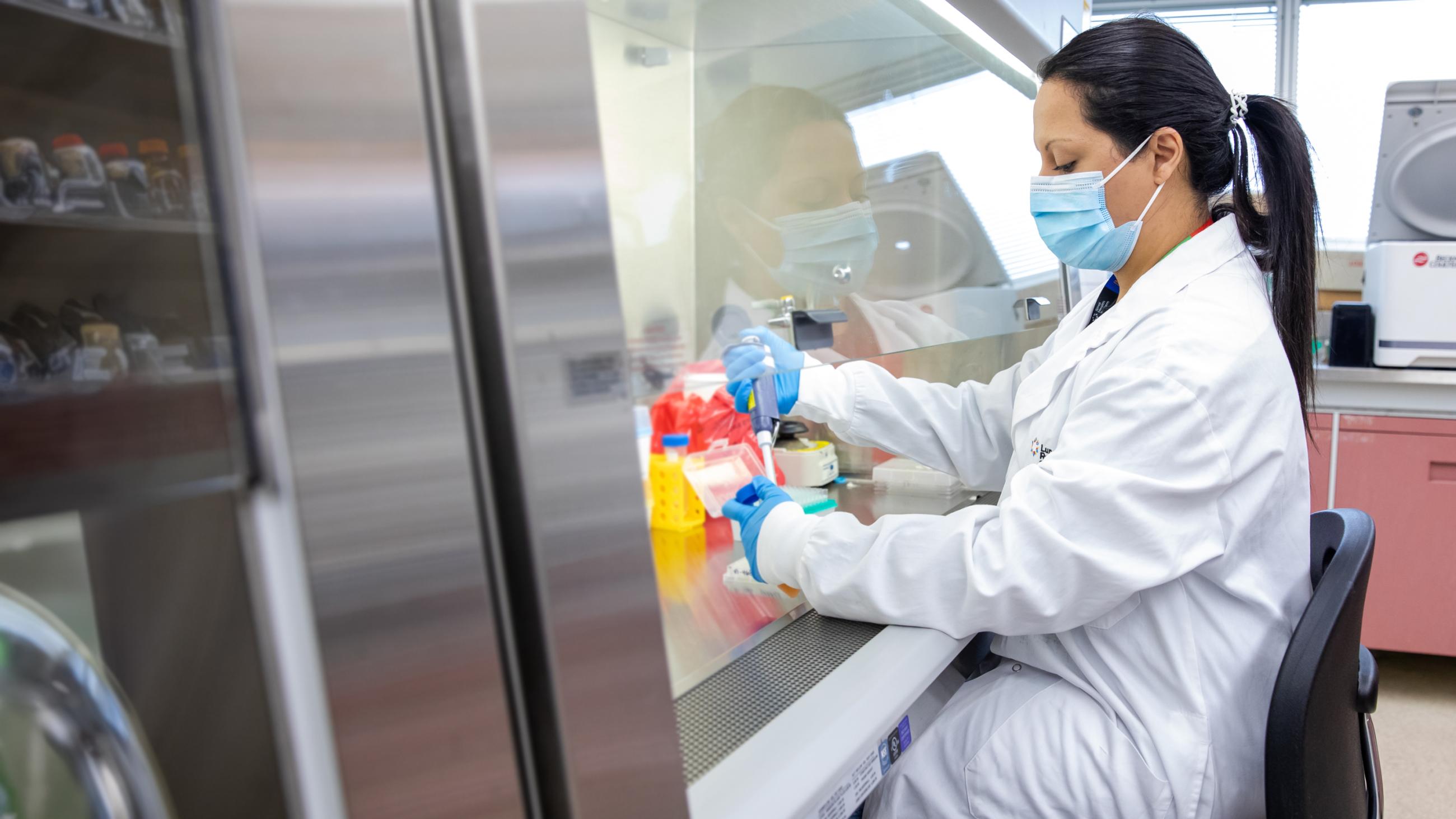
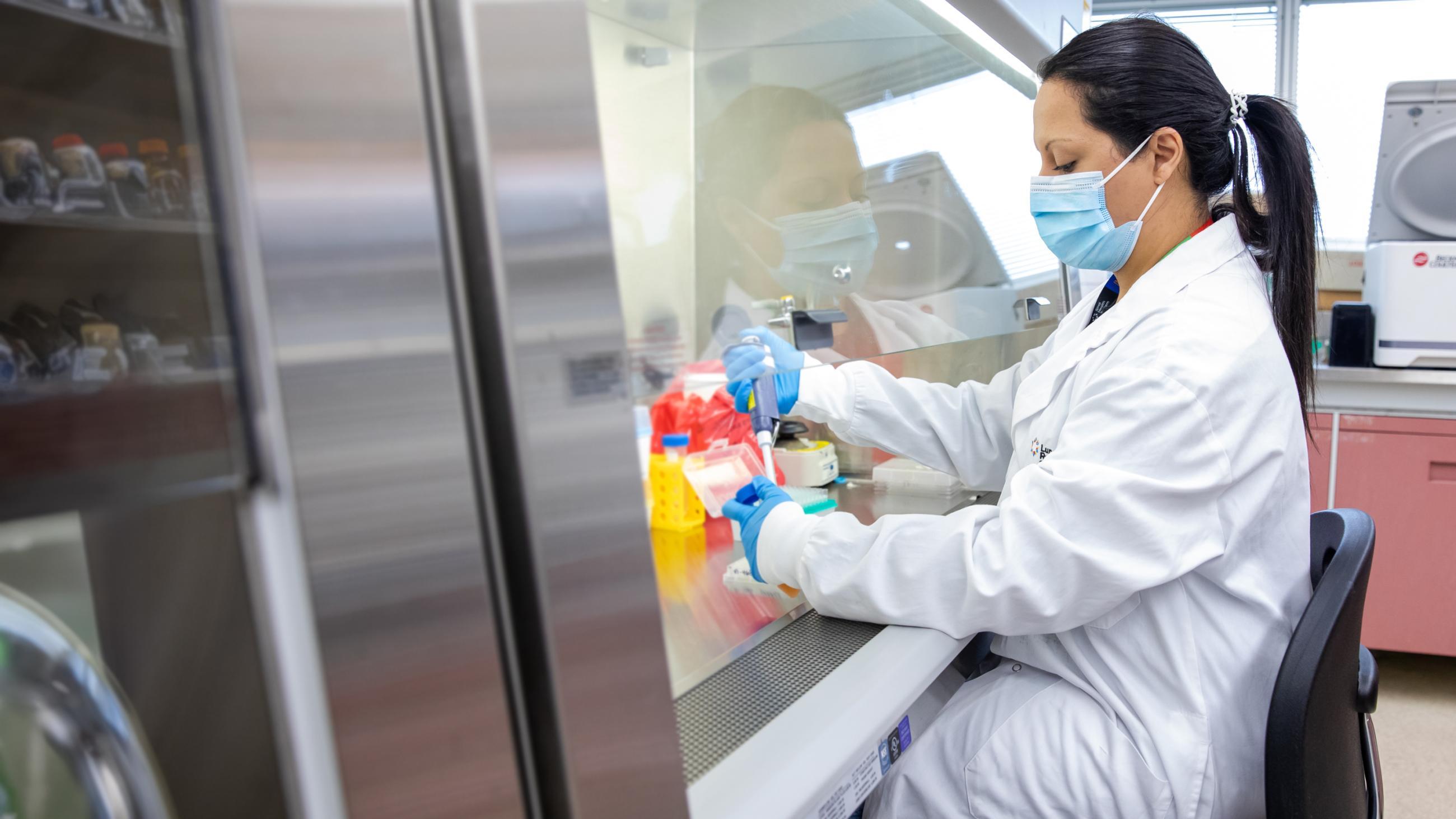
The new study, out today in the journal Nature, was conducted in the laboratory of Dr. Anne-Claude Gingras, a senior investigator at the Lunenfeld-Tanenbaum Research Institute (LTRI) and professor in the Department of Molecular Genetics at the University of Toronto.
Co-first authors Christopher Go and Dr. James Knight surveyed the human cell landscape using 192 markers for proteins known to reside in specific organelles that can “tag” neighbouring proteins in the same compartment.
“The Human Cell Map was able to predict the localization of 4,000 proteins across all compartments in living human cells,” said Go. “We sampled all the major organelles of the human cell and used innovative analysis to create the highest resolution map to date, with high accuracy in predicting novel localizations for many unmapped proteins.”
The human body is composed of trillions of cells that are each further separated into different compartments with dedicated functions, much like a house has different rooms for sleeping or preparing food. These compartments, called organelles, each contain different proteins that perform specific activities associated with the compartment. The mitochondria, the so-called powerhouse of the cell, is one example of an organelle.
The scientists said knowing which proteins are residing in which organelles is an important first step towards understanding the role of each cellular protein. Earlier approaches often used methods that first kill the cells before trying to separate the organelles from one another.
“Previously, it was like taking the house apart to isolate each of the individual rooms,” said Go. “These approaches tend to provide only crude views of the organization of a cell.”
Given the expansive nature of the Human Cell Map, the team also created an analysis portal to allow researchers around the world to delve deeper into the data. Users can scan each of the 192 markers in detail and compare their own data on protein localization to predictions made in the Human Cell Map.
Knight said while this work provides a greater understanding of the organization within the human cell, it also can be leveraged to better understand what happens during disease.
“Human diseases are typically characterized at the molecular level by proteins with aberrant behaviour that cause the cell to behave in pathological ways. In these situations, proteins will often change where they reside in the cell,” said Dr. Knight. “Our research is a first step in addressing this challenge in normal cells and we can use it for comparisons against altered cell states, such as disease conditions, to identify proteins with unexpected localizations that may help us understand a diseased cell better.”
The team said the map will now be used in a variety of projects to help shed additional light on protein localization in human cells. Future efforts will include using chemical, viral and disease conditions to better characterize how cells adapt structurally to these stressors. This can inform future research efforts towards a mechanistic understanding of diseased states and the development of future therapeutics.
This work was made possible by a Canadian Institutes of Health Research grant, as well as funding to the Network Biology Collaborative Centre at the LTRI, a facility supported by the Canada Foundation for Innovation funding, the Ontario Government, Genome Canada and Ontario Genomics. Funding for the analysis portal was provided by Compute Canada.
Story written by Francesco Zangari